此條目
無法為不熟悉主題的讀者提供充分的背景信息 。
(2023年1月1日 ) 請協助提供 更多背景信息。
候選門級輻射類群
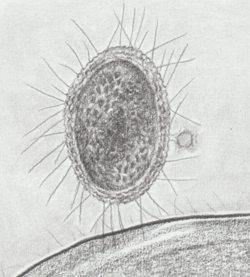
Drawing of a CPR bacterium from a "GWB1" sample.
科學分類
域:
細菌域 Bacteria
(未分級) :地位未定 incertae sedis
(未分級) :細菌候選門 Bacteria candidate phyla
下界:
候選門級輻射類群 Candidate phyla radiation
異名
候選門級輻射類群 (英語:candidate phyla radiation ,簡稱CPR group 細菌 的一個演化輻射 分支,其中的物種大多數尚未被培養,只能從宏基因組 和單細胞測序 得知其存在。由於它們的體積只有納米級,與其他細菌相比極為微小,因而也被稱為「納米細菌」(英語:nanobacteria ,納米細菌 也可以指一類曾被認為是細菌的納米級大小的礦物,二者沒有直接關係)或「超小細菌」(英語:ultra-small bacteria )。
在研究早期(2016年前後),候選門級輻射類群被認為可能含有70多個門 ,代表了細菌中15%的門級多樣性,[ 1] 基因組分類資料庫 [ 2] [ 3] 基因組 較小,缺少幾種生物合成途徑以及核糖體 蛋白。因此人們猜測它是專性共生 生物。[ 4] [ 5]
早期研究中,學者將現屬於本類群的幾個門歸類為一個超門,命名為Patescibacteria ,[ 6] 異名 ,[ 7] [ 2]
儘管有一些例外,本類群的成員基本上缺少一些胺基酸 和核苷酸 的生物合成 通路。目前為止,沒有基因組證據可以證明它們能產生一些細胞套膜 [ 5] 三羧酸循環 和電子傳遞鏈 複合體 ,包括三磷酸腺苷合酶 。這些通路在大多數自由生活的原核生物中廣泛存在,該類群缺少這些通路可能說明該類群可能主要由專性發酵共生生物組成。[ 8]
本類群的成員一般難以培養 ,因此在依賴培養的研究方法中會被遺漏。此外,它們有一些獨特的核糖體特徵,因此在依賴16S rRNA 的研究中也會被遺漏。它們的rRNA基因似乎負責編碼蛋白質,並且含有自我剪接內含子 [ 9] 16S rRNA 的研究中無法被檢測到;此外,所有成員都缺乏L30核糖體蛋白質 [ 8]
它們的許多特徵與DPANN 古菌 相似或類似。[ 5]
2016年的一個基於核糖體蛋白的生命之樹。[ 4] 基於核糖體蛋白和RNA聚合酶亞基的細菌和古細菌的系統發育[ 10] 早期的一些基於核糖體蛋白和蛋白家族 的系統發育研究認為候選門級輻射類群是細菌中最早分化的譜系,以下是各個門(正常字體)和超門(粗體)之間的系統發育關係:[ 4] [ 5]
然而,一些最近的研究認為,候選門級輻射類群屬於大地菌 類,與綠彎菌門 關係較近,[ 10] [ 11] [ 12]
細菌域 (Bacteria)
薄壁菌門 (Gracilicutes)
大地菌 (Terrabacteria)
根據{{tsl|en|International Code of Nomenclature of Prokaryotes|國際原核生物命名規約]],原核生物類群必須有純培養物 才能獲得正式名稱,而候選門級輻射類群的絕大多數物種無法滿足這個資格。然而,一些臨時名稱或候選名稱 [ 6] [ 13] [ 1]
Microgenomatia的系統發育[ 14] [ 15] [ 16]
"Woykebacterales" (CG2-30-54-11)
"Curtissbacterales"
"Daviesbacterales"
"Roizmanbacterales" (UBA1406)
"Gottesmanbacterales" (UBA10105)
"Levybacterales"
GWA2‑44‑7
"Amesbacteraceae"
"Blackburnbacteraceae" (UBA10165)
"Woesebacteraceae" (UBA8517)
"Shapirobacterales" (UBA12405)
"Chazhemtobacteriales"
"Beckwithbacteraceae" (CG1-02-47-37)
"Collierbacteraceae" (UBA12108)
"Chazhemtonibacteraceae"
"Chisholmbacteraceae"
"Cerribacteraceae" (UBA12028)
"Pacebacteraceae" (PJMF01)
Gracilibacteria的系統發育[ 14] [ 15] [ 16]
"Absconditabacteria"
"Ca. Altimarinus " {BD1-5: UBA6164}
"Absconditicoccaceae" {"Absconditabacterales"}
"Gracilibacteria"
"Abawacabacteriales" (RBG-16-42-10)
"Peregrinibacterales" (UBA1369)
"Fallacibacteriales" (UBA4473)
"Peribacterales"
ABY1的系統發育[ 14] [ 15] [ 16]
"Kerfeldbacterales" (SBBC01)
"Jacksonbacterales" (UBA9629)
"Komeilibacterales" (UBA1558)
"Kuenenbacterales" (UBA2196)
"Veblenbacterales"
"Magasanikbacterales"
"Uhrbacterales" (SG8-24)
"Buchananbacterales"
"Falkowbacterales" (BM507)
"Moisslbacterales" (UBA2591)
Paceibacteria的系統發育[ 14] [ 15] [ 16]
"Moranbacterales"
UBA6257
"Brennerbacteraceae"
"Jorgensenbacteraceae" (GWB1-50-10)
"Wolfebacteraceae" (UBA9933)
"Colwellbacteraceae" (UBA9933)
"Harrisonbacteraceae" (WO2-44-18)
"Liptonbacteraceae" (2-01-FULL-56-20)
"Spechtbacterales"
"Terrybacterales"
"Parcunitrobacterales" (GWA2-38-13b)
"Portnoybacterales"
"Paceibacterales"
"Wildermuthbacteraceae" (UBA10102)
"Gribaldobacteraceae" (CG1-02-41-26)
"Paceibacteraceae" ("Parcubacteria")
"Nealsonbacteraceae" (PWPS01)
"Staskawiczbacteraceae"
"Azambacterales" (UBA10092)
"Yanofskybacterales" (2-02-FULL-40-12)
"Sungbacterales"
"Ryanbacterales"
"Giovannonibacterales" (UBA11713)
"Niyogibacterales" (HO2-45-28)
"Tagabacterales"
UBA9983
"Vogelbacteraceae" (XYD1-FULL-46-19)
"Yonathbacteraceae" (UBA1539)
"Nomurabacteraceae" (UBA9973)
"Adlerbacteraceae" (SBAW01)
"Kaiserbacteraceae" (UBA2163)
"Campbellbacteraceae" (UBA12079)
"Taylorbacteraceae" (PALSA-1337)
"Zambryskibacteraceae"
?"Elulimicrobiota " Rodriguez-R et al. 2020
Clade "Patescibacteria" Rinke et al. 2013
"Wirthbacteria " Hug et al. 2016
Microgenomates Cluster
Gracilibacteria Cluster
Clade "Absconditabacteria"
Superphylum "Gracilibacteria"
Saccharibacteria Cluster
Parcubacteria Cluster
目前的系統發育結果基於核糖體蛋白 。[ 4] 蛋白家族 和16S 核糖體RNA ,得到的結果類似但解析度較低。[ 1] [ 17]
^ 1.0 1.1 1.2 Danczak RE, Johnston MD, Kenah C, Slattery M, Wrighton KC, Wilkins MJ. Members of the candidate phyla radiation are functionally differentiated by carbon- and nitrogen-cycling capabilities . Microbiome. September 2017, 5 (1): 112. PMC 5581439 PMID 28865481 doi:10.1186/s40168-017-0331-1 ^ 2.0 2.1 Parks, Donovan; Chuvochina, Maria; Waite, David; Rinke, Christian; Skarshewski, Adam; Chaumeil, Pierre-Alain; Hugenholtz, Philip. A standardized bacterial taxonomy based on genome phylogeny substantially revises the tree of life . Nature Biotechnology. 27 August 2018, 36 (10): 996–1004 [13 January 2021] . PMID 30148503 S2CID 52093100 doi:10.1038/nbt.4229 ^ Parks, Donovan H.; Rinke, Christian; Chuvochina, Maria; Chaumeil, Pierre-Alain; Woodcroft, Ben J.; Evans, Paul N.; Hugenholtz, Philip; Tyson, Gene W. Recovery of nearly 8,000 metagenome-assembled genomes substantially expands the tree of life. Nature Microbiology. November 2017, 2 (11): 1533–1542. PMID 28894102 doi:10.1038/s41564-017-0012-7 ^ 4.0 4.1 4.2 4.3 Hug LA, Baker BJ, Anantharaman K, Brown CT, Probst AJ, Castelle CJ, et al. A new view of the tree of life. Nature Microbiology. April 2016, 1 (5): 16048. PMID 27572647 doi:10.1038/nmicrobiol.2016.48 ^ 5.0 5.1 5.2 5.3 Castelle CJ, Banfield JF. Major New Microbial Groups Expand Diversity and Alter our Understanding of the Tree of Life. Cell. March 2018, 172 (6): 1181–1197. PMID 29522741 doi:10.1016/j.cell.2018.02.016 ^ 6.0 6.1 Rinke C; et al. Insights into the phylogeny and coding potential of microbial dark matter. Nature. 2013, 499 (7459): 431–7. Bibcode:2013Natur.499..431R PMID 23851394 doi:10.1038/nature12352 hdl:10453/27467 ^ Beam, Jacob P.; Becraft, Eric D.; Brown, Julia M.; Schulz, Frederik; Jarett, Jessica K.; Bezuidt, Oliver; Poulton, Nicole J.; Clark, Kayla; Dunfield, Peter F.; Ravin, Nikolai V.; Spear, John R.; Hedlund, Brian P.; Kormas, Konstantinos A.; Sievert, Stefan M.; Elshahed, Mostafa S.; Barton, Hazel A.; Stott, Matthew B.; Eisen, Jonathan A.; Moser, Duane P.; Onstott, Tullis C.; Woyke, Tanja; Stepanauskas, Ramunas. Ancestral Absence of Electron Transport Chains in Patescibacteria and DPANN . Frontiers in Microbiology. 2020, 11 : 1848. PMC 7507113 PMID 33013724 doi:10.3389/fmicb.2020.01848 ^ 8.0 8.1 Brown CT, Hug LA, Thomas BC, Sharon I, Castelle CJ, Singh A, et al. Unusual biology across a group comprising more than 15% of domain Bacteria (PDF) . Nature. July 2015, 523 (7559): 208–11. Bibcode:2015Natur.523..208B OSTI 1512215 PMID 26083755 S2CID 4397558 doi:10.1038/nature14486 ^ Belfort M , Reaban ME, Coetzee T, Dalgaard JZ. Prokaryotic introns and inteins: a panoply of form and function . Journal of Bacteriology. July 1995, 177 (14): 3897–903. PMC 177115 PMID 7608058 doi:10.1128/jb.177.14.3897-3903.1995 ^ 10.0 10.1 Martinez-Gutierrez CA, Aylward FO. Phylogenetic signal, congruence, and uncertainty across bacteria and archaea . Molecular Biology and Evolution. 2021, 38 (12): 5514–5527. PMC 8662615 PMID 34436605 doi:10.1093/molbev/msab254 ^ Coleman GA, Davín AA, Mahendrarajah TA, Szánthó LL, Spang A, Hugenholtz P, Szöllősi GJ, Williams TA. A rooted phylogeny resolves early bacterial evolution . Science. 2021, 372 (6542). PMID 33958449 S2CID 233872903 doi:10.1126/science.abe0511 hdl:1983/51e9e402-36b7-47a6-91de-32b8cf7320d2 ^ Taib N, Megrian D, Witwinowski J, Adam P, Poppleton D, Borrel G, Beloin C, Gribaldo S. Genome-wide analysis of the Firmicutes illuminates the diderm/monoderm transition (PDF) . Nature Ecology and Evolution. 2020, 4 (12): 1661–1672. PMID 33077930 S2CID 224810982 doi:10.1038/s41559-020-01299-7 ^ Sayers. Patescibacteria group . National Center for Biotechnology Information (NCBI) taxonomy database. [2021-03-20 ] . ^ 14.0 14.1 14.2 14.3 14.4 GTDB release 09-RS220 . Genome Taxonomy Database . [10 May 2024] . ^ 15.0 15.1 15.2 15.3 15.4 bac120_r220.sp_labels . Genome Taxonomy Database . [10 May 2024] . ^ 16.0 16.1 16.2 16.3 16.4 Taxon History . Genome Taxonomy Database . [10 May 2024] . ^ Méheust, Raphaël; Burstein, David; Castelle, Cindy J.; Banfield, Jillian F. The distinction of CPR bacteria from other bacteria based on protein family content . Nature Communications. 13 September 2019, 10 (1): 4173. Bibcode:2019NatCo..10.4173M PMC 6744442 PMID 31519891 doi:10.1038/s41467-019-12171-z
Most of the Tree of Life is a Complete Mystery . We know certain branches exist, but we have never seen the organisms that perch there. by Ed Yong, April 12, 2016, atlantic.com.Ultra-Small, Parasitic Bacteria Found in Groundwater, Dogs, Cats — And You ; on: SciTechDaily; July 21, 2020; source: Forsyth InstituteMcLean, Jeffrey S.; Bor, Batbileg; Kerns, Kristopher A.; Liu, Quanhui; To, Thao T.; Solden, Lindsey; Hendrickson, Erik L.; Wrighton, Kelly; Shi, Wenyuan; He, Xuesong. Acquisition and Adaptation of Ultra-small Parasitic Reduced Genome Bacteria to Mammalian Hosts . Cell Reports. 2020, 32 (3): 107939. PMC 7427843 PMID 32698001 bioRxiv 10.1101/258137 doi:10.1016/j.celrep.2020.107939 Bokhari, RH; Amirjan, N; Jeong, H; Kim, KM; Caetano-Anollés, G; Nasir, A. Bacterial Origin and Reductive Evolution of the CPR Group . Genome Biology and Evolution. 1 March 2020, 12 (3): 103–121. PMC 7093835 PMID 32031619 doi:10.1093/gbe/evaa024


 . PMID 28865481. doi:10.1186/s40168-017-0331-1
. PMID 28865481. doi:10.1186/s40168-017-0331-1  .
.
 .
.
 .
.
 .
.
 . hdl:10453/27467
. hdl:10453/27467  .
.
 . PMID 33013724. doi:10.3389/fmicb.2020.01848
. PMID 33013724. doi:10.3389/fmicb.2020.01848  .
.
 . PMID 7608058. doi:10.1128/jb.177.14.3897-3903.1995.
. PMID 7608058. doi:10.1128/jb.177.14.3897-3903.1995.
 . PMID 34436605. doi:10.1093/molbev/msab254.
. PMID 34436605. doi:10.1093/molbev/msab254.
 .
.
 . PMID 31519891. doi:10.1038/s41467-019-12171-z
. PMID 31519891. doi:10.1038/s41467-019-12171-z  .
.
 . PMID 32698001. bioRxiv 10.1101/258137
. PMID 32698001. bioRxiv 10.1101/258137  . doi:10.1016/j.celrep.2020.107939.
. doi:10.1016/j.celrep.2020.107939. . PMID 32031619. doi:10.1093/gbe/evaa024
. PMID 32031619. doi:10.1093/gbe/evaa024  .
.